Haonan Wu (吴昊楠)
Department of Computer Science and Engineering, The Pennsylvania State University.

506, Wartik Lab
360 Science Drive
State College, PA, USA
Hi guys! I am a Ph.D. Candidate in Computer Science and Engineering at Penn State, under the supervision of Dr. Paul Medvedev.
Previously, I obtained my M.S. in Operations Research (OR) and B.S. in Biotechnology at Shandong University.
Research interests
I am interested in developing theories, algorithms, and practical tools for bioinformatics, with the goal of addressing biological questions. My research is guided by two principles:
- Developing methods grounded in rigorous mathematics and algorithms, or in clear mathematical intuition.
- Building practically useful tools that incorporate effective heuristics and benefit the bioinformatics community.
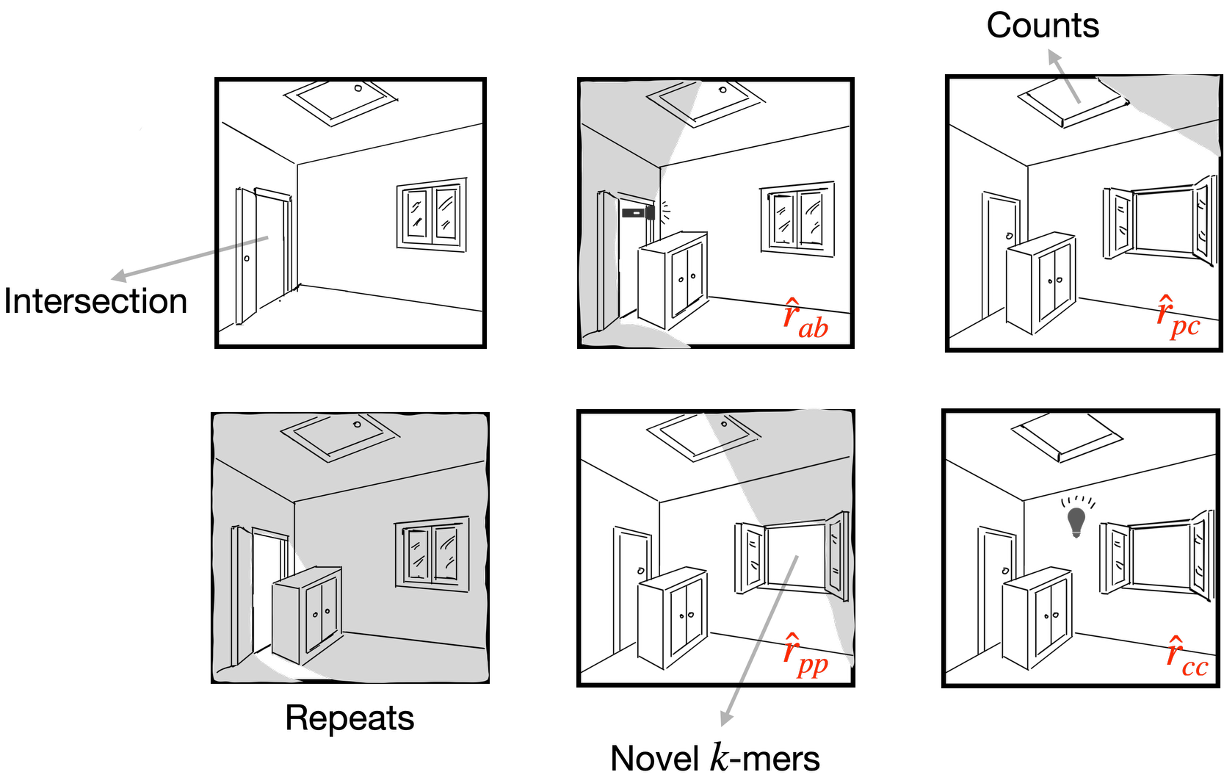
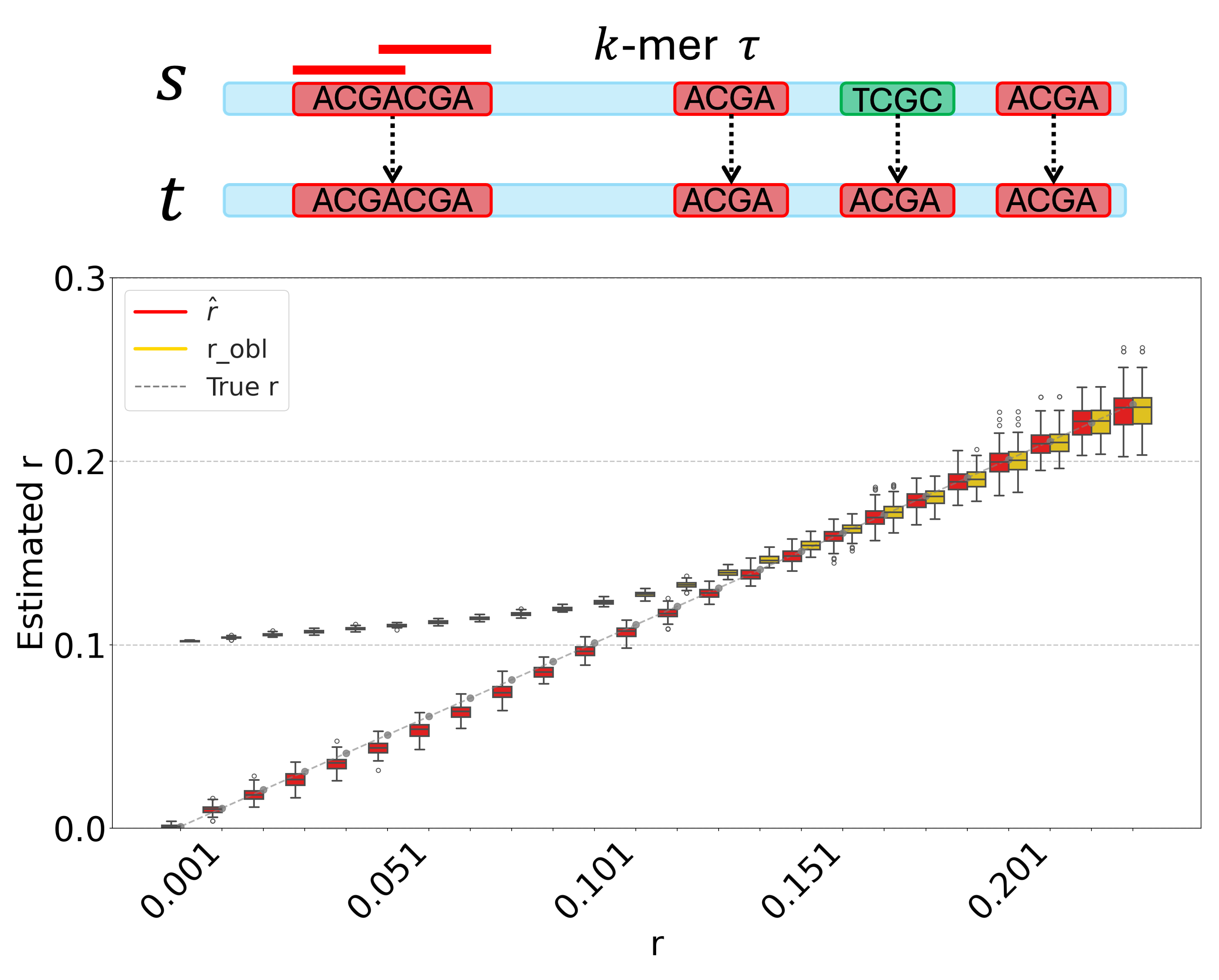
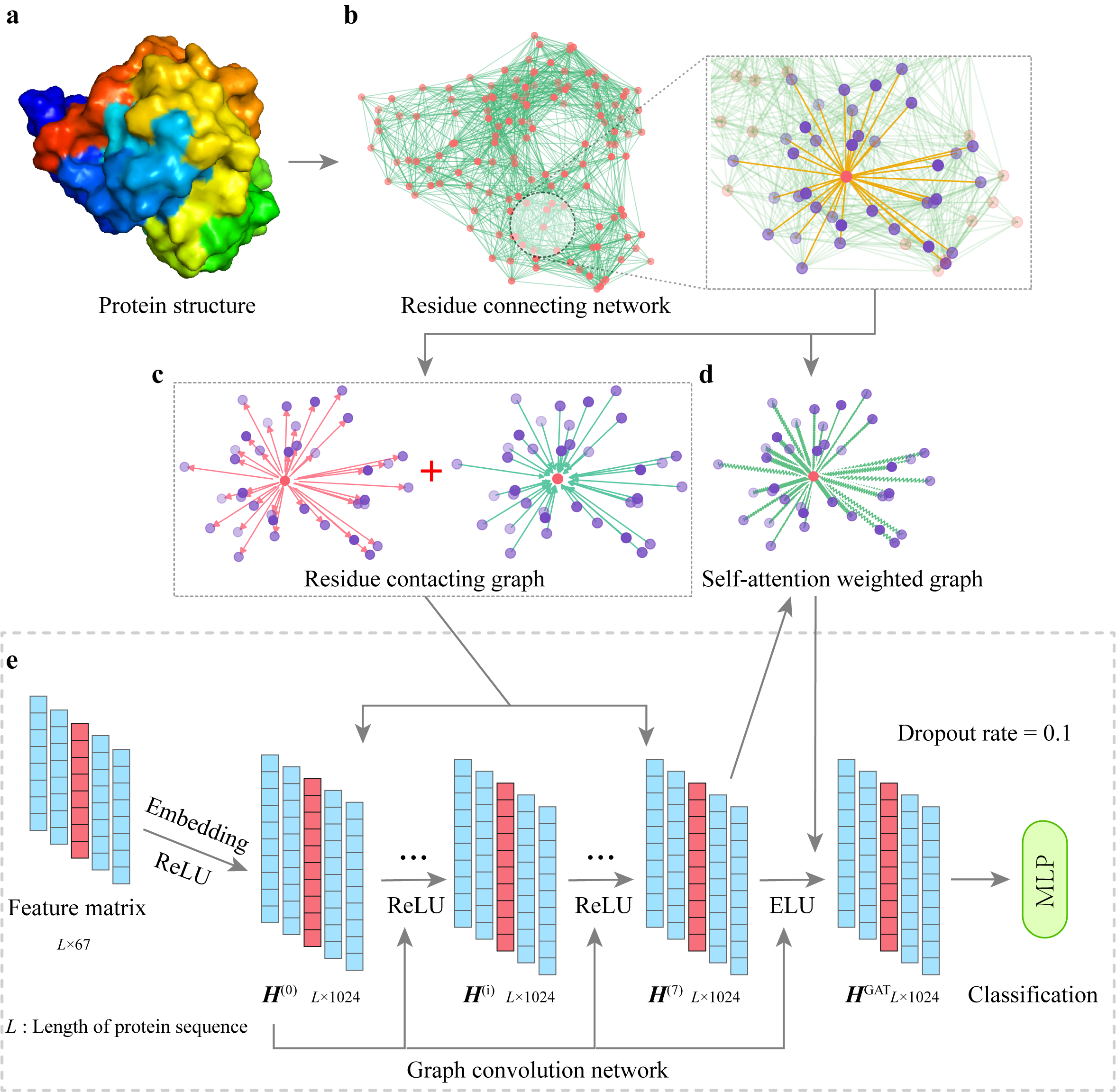
Currently, my research focuses on repetitive genomic sequences from both theoretical and practical perspectives. My theoretical work focuses on the analysis of $k$-mer behavior under mutation and evolutionary processes, and estimation of the mutation rate. On the practical side, I develop algorithms specifically designed for centromeres, covering tasks such as annotation, variant calling, and sequence alignment.
news
| Jun 01, 2026 | I was selected for an ISMB 2026 Conference Fellowship. 🎉 |
|---|---|
| Apr 24, 2026 | I passed the Comprehensive Exam in the CSE PhD program at Penn State. 🍾 |
| Apr 02, 2026 | Our paper “The gift of novelty: repeat-robust k-mer-based estimators of mutation rates” was accepted by ISMB2026. 🎉 |
| Nov 06, 2025 | I give a talk at Genome Informatics 2025 with title “Don’t repeat no repeat: A k-mer-based estimator of the substitution rate between repetitive sequences”. |
| Oct 08, 2025 | I give the talk “Don’t repeat no repeat: A k-mer-based estimator of the substitution rate between repetitive sequences” at WABI2025 |