Momonmer Types, Suprachromosomal Families (SFs) and HORs Identification
In this blog, we will introduce the structure/concepts in centromere study, like monomer types, Suprachromosomal Families (SFs), HORs. What is the modern understanding of these concepts. Then, we will reveiw how biologist studied these structure? In other words, how people identifies these ground truth in centromere.
Hierarchies in Alpha-Satellite Array
Model of Centromeric Regions
Alpha satellite DNAs are credited as the genetic substrate of endogenous centromeres in primates, starting with the new-world monkeys. No alpha satellites have been found in tarsiers and lemurs 12. In humans, arrays of alpha satellites are organized in discrete layers expanding out from a multimegabase-sized homogeneous core that is composed of chromosome-specific HORs (live or active arrays). Additional subsets of alpha satellites are often observed on one or both sides of the core in a near symmetrical formation. From center to outside, we have
-
A zone of smaller homogeneous HOR arrays (pseudocentromeres or inactive arrays)
-
An outermost layer of progressively more divergent and smaller (center-to-periphery gradient) HOR
-
Monomeric arrays (relic centromeres)
Both inactive HOR arrays and divergent arrays are often in the range of a few to hundreds of kilobases. Other distinct satellite classes, such as the classical human satellites (human satellites 1–3, or HSat1–HSat3), are of variable size (up to several megabases) and positioned in the adjacent pericentromeric regions. Segmental duplications are often observed directly flanking the satellite arrays or in centromeric transition regions extending out to the p-arm or q-arm (greater than a megabase) or between adjacent satellite arrays. The entire centromeric region can be defined by those sequences in linkage or sharing a common centromere-spanning haplotype (cenhap), which is characterized by repressed meiotic recombination3.
Centromere expansion likely goes in waves of interchromosomal transfer and amplification, where the HORs (or monomeric sequences) of the newly formed novel centromere jump from one live centromere to another and amplify in the new location to form the next generation of live centromeres (a centromeric layer) in all chromosomes or in a group of chromosomes. The remnants of the old centromere are displaced sideways, shrink, diverge, and structurally degrade.
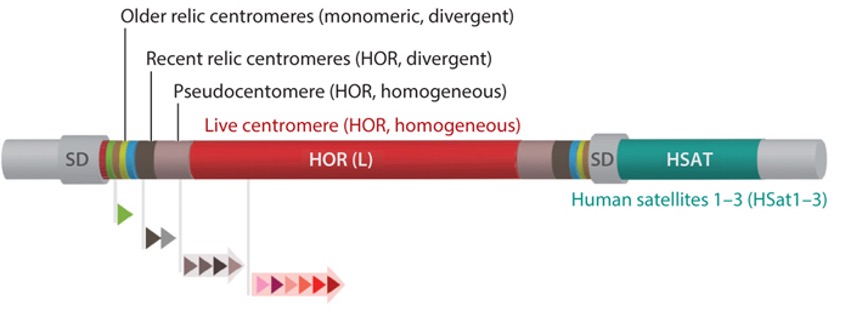
Fig 1. General genomic structure of a human centromeric region, which includes one homogeneous core made of chromosome-specific HORs (red) and the imperfect symmetrical organization of smaller arrays of various other homogeneous HORs [pseudocentromeres or inactive HOR arrays (light gray)], divergent HORs [recent relic centromeres (dark gray)], and multiple distinct divergent monomeric arrays (older relic centromeres). These regions typically include other pericentromeric satellite classes [e.g., HSat1–HSat3 (teal)] and SDs. The entire centromeric region is defined by those sequences in the cenhap, presented as gray flanking regions extending into the p-arm and q-arm. Arrayed triangles indicate alpha satellite monomers and HORs of various length and structures composed of several different monomers.
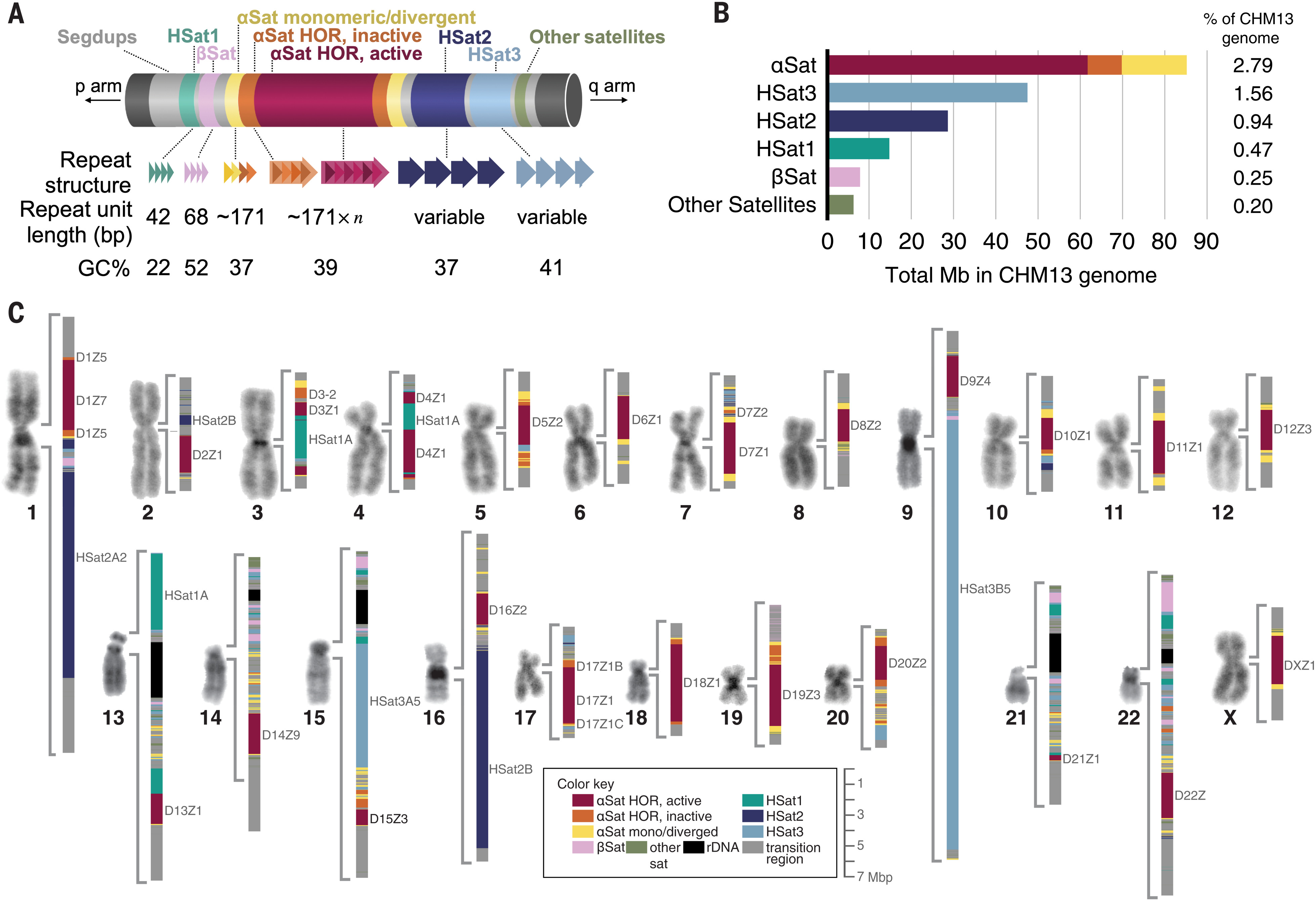
Fig 2. Overview of all peri/centromeric regions in CHM13.
Monomer Class and Monomer Type
The individual monomers within a HOR unit have 50–70% identity and can be distinguished such that HOR unit length is determined by where the next monomer shows nearly total sequence identity to the first monomer in the HOR.
SF and A/B type
HOR and naming system
Where does the ground truth come from
Reference
-
Miga, Karen H., and Ivan A. Alexandrov. “Variation and evolution of human centromeres: a field guide and perspective.” Annual review of genetics 55.1 (2021): 583-602. ↩
-
The Evolutionary Origin of Man Can Be Traced in the Layers of Defunct Ancestral Alpha Satellites Flanking the Active Centromeres of Human Chromosomes. ↩
-
Langley, Sasha A., et al. “Haplotypes spanning centromeric regions reveal persistence of large blocks of archaic DNA.” elife 8 (2019): e42989. ↩
Enjoy Reading This Article?
Here are some more articles you might like to read next: