Summary for Latest studies on Centromere Evolution and Mutation
The kinetochore assembles at the most homogeneous subregion of the active HOR array
Human centromeres are organized as megabase-scale arrays of alpha-satellite higher-order repeats (HORs). Centromeres can contain multiple distinct alpha satellite HOR arrays that can be classified into active and inactive HORs. Centromere protein A (CENP-A) is an H3 variant that is enriched in centromeric nucleosomes and marks sites of kinetochore assembly. Centromeres are typically hypermethylated throughout the active HOR array. But in some small regions, snotable hypomethylation is colocalizing with CENP-A enrichment and we refer this hypomethylation as the centromeric dip region (CDR) (Fig. 2). CDRs are present only in active HORs, and that active HORs were larger in size and had higher mean methylation frequency than inactive HORs, as exemplified by the chromosome 5 centromere (Fig. 1). The CDR and CENP-A occupancy are tightly correlated and together define the functional kinetochore site 1.
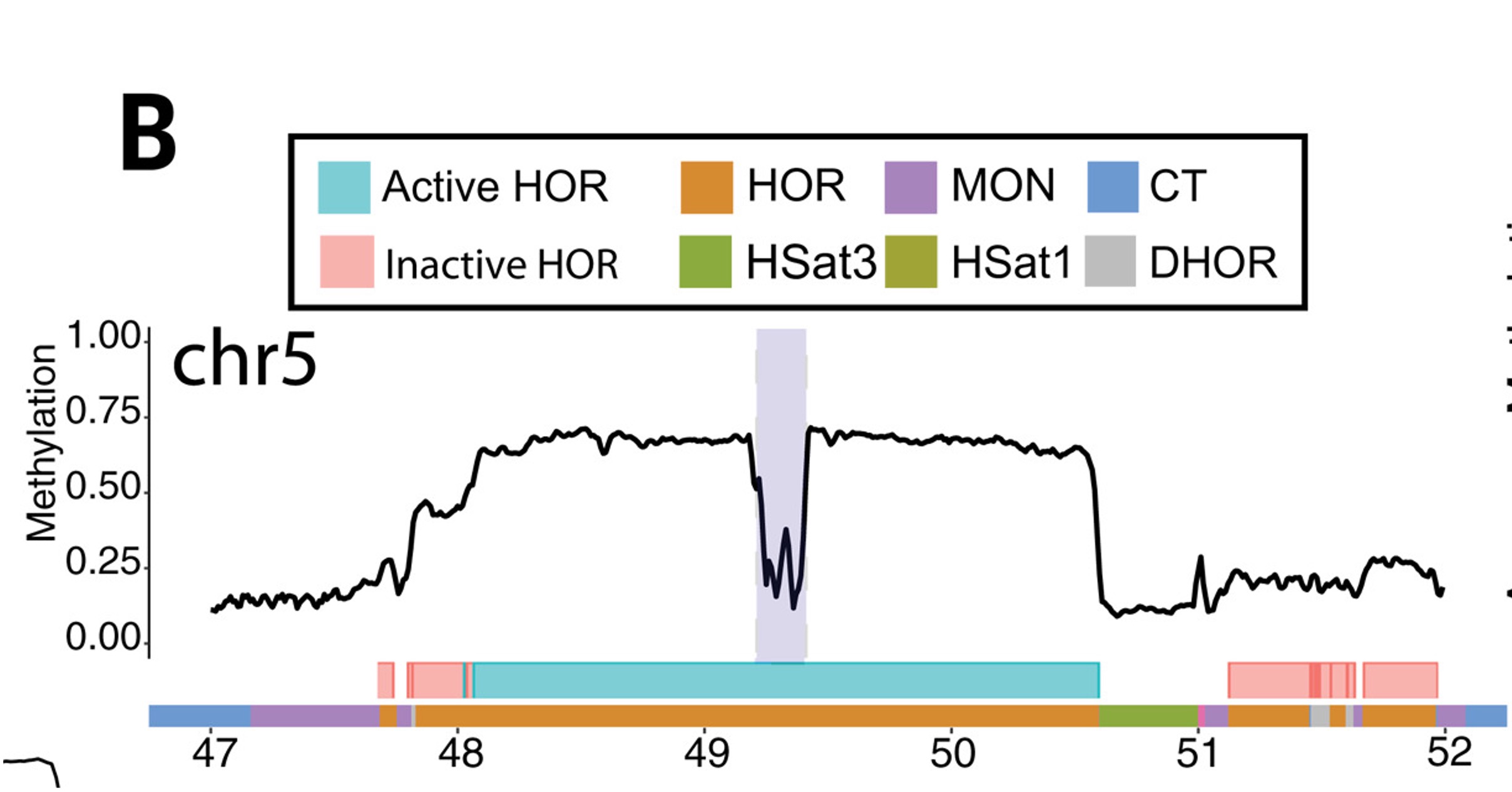
Fig.1 CHM13 methylation in the centromeric region of chromosome 5. Smoothed methylation frequency is plotted in 10-kb bins. HOR arrays are annotated as blue (“active”) and pink (“inactive”).1.

Fig.2 Smoothed methylation frequency in 10-kb bins of the active HOR array for all CHM13 chromosomes. CENP-A enrichment from CUT&RUN data is shown as a heatmap under each plot. Chromosomes 3 and 4 have an HSat1 repeat (blue highlight) that breaks up the live HOR array.1.
Miga & Alexandrov 2 found that an average divergence of neighboring copies of a repeat in an array, as there is a gradient of intra-array divergence from the center to the periphery that reflects the age of alpha satellite arrays. Altemose et al. 3 found that the central part of the active array contain HOR variants slightly different from those on the flanks. They aligned individual HOR units within the same array and clustered them on the basis of their shared sequence variants into “HOR-haplotypes” or “HOR-haps”. Then confirmed that the middle HOR-haps are the most recently evolved. Then, they observed that human centromeres and CDRs are typically, although not universally, positioned over young and/or recently expanded layers within active HOR arrays in CHM13, indicating that centromere function is closely related to the rapid evolution of alpah-Sat sequences. Gao et al.4 show that the putative kinetochore site, marked by the CDR and an enrichment of CENP-A chromatin, typically resides within the most highly identical and homogenous sequences of the centromeric α-satellite HOR array. (1. mean sequence identity within the CDR and across the α-satellite HOR array are 99.55% vs. 99.36%, respectively; p<0.0001 2. CDR resides within regions that are more homogenous in sequence and structure and, therefore, have lower entropy (mean entropy = 0.74 vs. 0.80, respectively; p<0.0001)) So, I tentatively claim that most recent expanded HOR has the least intra-array divergence, so CDRs are co-localized in this region. The exceptions observed from Altemose et al. that some CDRs are not located in young layers, because some recent expanded HOR is not the least intra-array divergence regions.
High sequence homogeneity elevates mutation rate
Gao et al.4 leveraged genome assemblies from a four-generation pedigree that had been recently generated to validate centromere mutation rates. The lowest mutation rate occurs in the unique sequences in the pericentromere (7.1×10-8 Ms/bp/G), while the monomeric/divergent α-satellite HORs have a slightly higher mutation rate (1.1×10-7 Ms/bp/G), and the active α-satellite HORs have a mutation rate 2.6×10-7 Ms/bp/G. Within the active α-satellite HOR array itself, we find that the CDR has the highest mutation rate (5.1×10-7 Ms/bp/G). For de novo SVs, they believe that elevated SV mutation rate in the CDR may reflect the higher sequence identity of these regions, which are more prone to homologous recombination, leading to expansions and deletions, relative to less identical sequences.
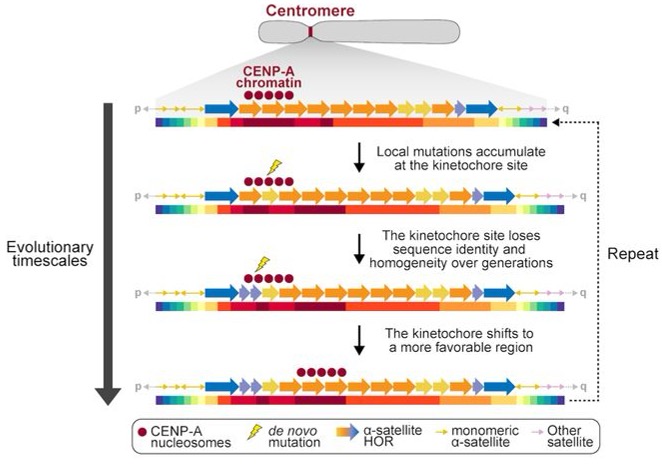
Fig.3 Model of centromere mutation and evolution, showing that the CDR is often located within the most identical and homogenous sequences of the centromere and is prone to both SNVs and SVs that alter the genetic and epigenetic landscape of the centromere over time.4.
Structural mutations remodel both the genetic and epigenetic landscape
Variation within α-satellite organization negatively affects centromere assembly and function. If variant and wild-type HORs are interspersed across the entire array, the irregularity might disrupt structural requirements for kinetochore architecture, such as CENP-C-mediated bridging between nucleosomes5.
At the epigenetic level, younger αSats expand in copy number within NativeCen, termed layered expansions, it is possible that the CDR shifts with these expansions during the normal process of HOR evolution, with the CpG hypermethylation boundary sliding along with it, but only within an array 6.
In extreme cases—when the most homogeneous region of the active array is disrupted beyond a functional threshold—the kinetochore may relocate entirely to a neighboring inactive array, a phenomenon documented on human chromosome 17 5.
Unified model: the complete evolutionary cycle
The centromere paradox is the evolutionary enigma where centromeric DNA sequences and their associated proteins evolve rapidly, yet their essential function—proper chromosome segregation during cell division—remains highly conserved 7.
To answer centromere paradox, Gao et al.’s 4findings provide the foundation for a unified model in which (i) kinetochores preferentially form on homogenous stretches of α-satellite HORs; (ii) those stretches are prone to elevated rates of homologous recombination and structural change because their sequence identity promotes misalignment and unequal exchange; and (iii) such genetic changes feed back on chromatin state and kinetochore localization, producing continual shifts in centromere architecture across generations (an evolutionary “treadmill” of DNA-protein interactions).
How to choose favorable HOR
1. Centromere Drive Hypothesis
While meiosis is symmetrical in males, resulting in four gametes, in most plants and animals, females undergo asymmetric meiosis, where only one of the resulting four meiotic products survives and develops into the oocyte, while the remaining three turn into polar bodies or degenerate. This asymmetry can result in competition between homologous chromosomes for inclusion in the oocyte (a phenomenon known as meiotic drive). According to the centromere-drive hypothesis, centromeric DNA acts as a selfish genetic element, exploiting asymmetric female meiosis to promote its preferential transmission to the egg 8. Lukáš Chmátal et al. 9 show this hypothesis in mice i.e. stronger centromeres, manifested by increased kinetochore protein levels and altered interactions with spindle microtubules, are preferentially retained in the egg.
2. Kinetochore Selection Hypothesis
(a) the evolution of centromeric repeats is not entirely neutral, and they are selected by the affinity to a kinetochore, which is free to move and chooses the most favorable place to reside within the live array;
(b) this selection operates through the ability of a kinetochore to amplify and possibly homogenize the repeats on which it resides (kinetochore-associated recombination machine, KARM) and
(c) the old centromere abandoned by the kinetochore degrades (deletions, inversions, TE insertions, HSat expansions, and hypermutability) 2.
Reference
-
Gershman, Ariel, et al. “Epigenetic patterns in a complete human genome.” Science 376.6588 (2022): eabj5089. ↩ ↩2 ↩3
-
Miga, Karen H., and Ivan A. Alexandrov. “Variation and evolution of human centromeres: a field guide and perspective.” Annual review of genetics 55.1 (2021): 583-602. ↩ ↩2
-
Altemose, Nicolas, et al. “Complete genomic and epigenetic maps of human centromeres.” Science 376.6588 (2022): eabl4178. ↩
-
Gao, Shenghan, et al. “A global view of human centromere variation and evolution.” bioRxiv (2025): 2025-12. ↩ ↩2 ↩3 ↩4
-
Sullivan, Lori L., Kimberline Chew, and Beth A. Sullivan. “α satellite DNA variation and function of the human centromere.” Nucleus 8.4 (2017): 331-339. ↩ ↩2
-
Hoyt, Savannah J., et al. “Haplotype-Resolved Genomics Reveals Conserved Chromatin Architecture and Epigenetic Constraints of Human Neocentromeres.” bioRxiv (2025): 2025-12. ↩
-
Henikoff, Steven, Kami Ahmad, and Harmit S. Malik. “The centromere paradox: stable inheritance with rapidly evolving DNA.” Science 293.5532 (2001): 1098-1102. ↩
-
Rosin, Leah F., and Barbara G. Mellone. “Centromeres drive a hard bargain.” Trends in Genetics 33.2 (2017): 101-117. ↩
-
Chmátal, Lukáš, et al. “Centromere strength provides the cell biological basis for meiotic drive and karyotype evolution in mice.” Current biology 24.19 (2014): 2295-2300. ↩
Enjoy Reading This Article?
Here are some more articles you might like to read next: